Our Research
|
Role of Ferritinophagy in Pancreatic Cancer
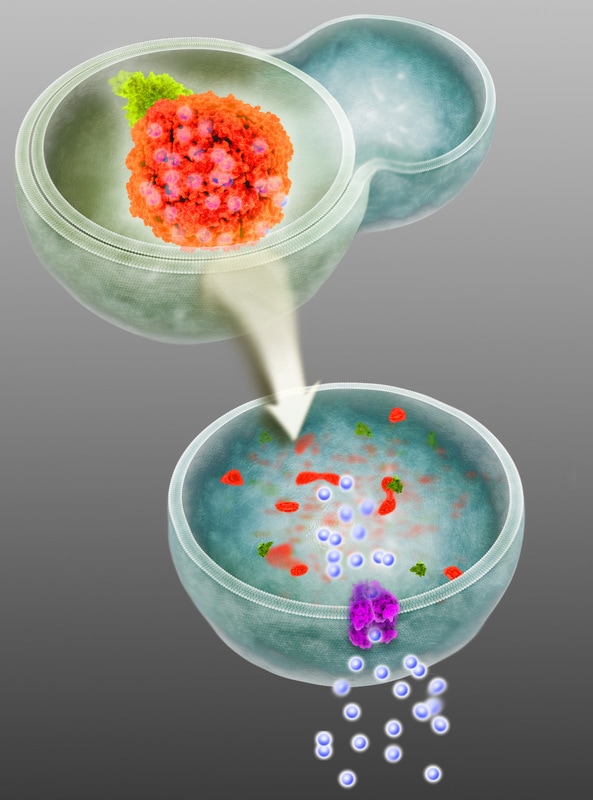
Recent work has shown that pancreatic cancers have a distinct dependence on autophagy (a cellular catabolic pathways) and that inhibition leads to DNA damage and metabolic defects leading to growth suppression. The unique importance of autophagy in pancreatic cancer suggests that there may be a subset of proteins that are specifically targeted by autophagy for degradation, which would promote proliferation and survival. To identify novel proteins degraded by autophagy in pancreatic cancer, we purified autophagosomes from pancreatic cancer cell lines and employed quantitative mass spectrometry-based proteomics. One of the most highly enriched autophagosomal proteins was NCOA4. Importantly, interaction proteomics revealed ferritin heavy and light chains (FTH1, FTL) as high confidence interacting proteins. Ferritin forms a complex of 24 subunits of light and heavy chains that is capable of storing up to 4,500 iron atoms. When iron levels in the cell are low, ferritin is degraded allowing the release of iron for use by the cell. Ferritin is degraded in the lysosome and recent evidence implicated autophagy in the process; however, the mechanism underlying this activity remained unclear. Our work identified NCOA4 as the selective autophagy receptor for ferritin autophagy (ferritinophagy) revealing the mechanism of how ferritin is degraded to regulate bioavailable iron levels (Mancias et al., Nature 2014) (Mancias et al., Elife 2015). Given the reliance of multiple cancers, including pancreatic cancer, on both high levels of autophagy as well as iron, understanding the role of NCOA4 in pancreatic cancer may reveal a new therapeutic approach. However, the role of ferritinophagy in the larger context of organismal iron metabolism may limit the therapeutic index of such a therapy; therefore, understanding the physiologic role of NCOA4 will be equally important. We will take an integrated biochemical, cell biologic, and in vivo approach in order to answer fundamental questions of the role of NCOA4-mediated ferritinophagy in pancreatic cancer and in systemic iron homeostasis. |
|
Selective Autophagy in Pancreatic Cancer
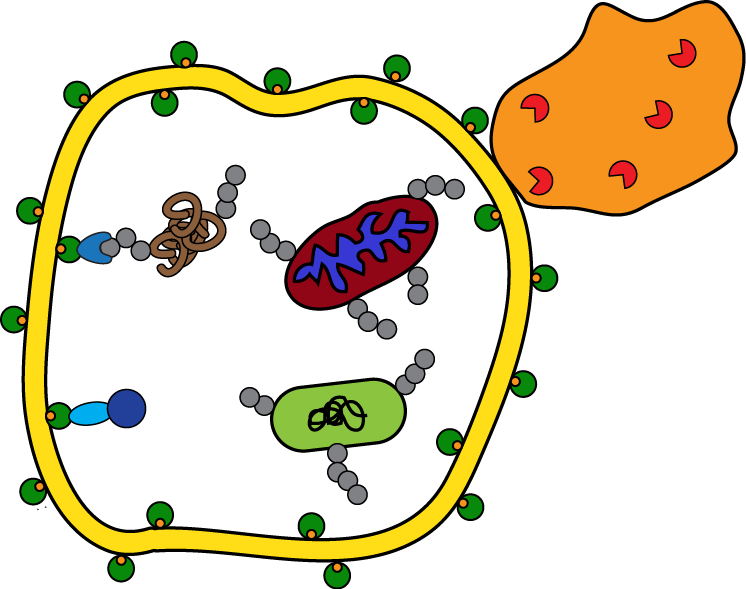
Using our quantitative autophagosomal proteomics methodology, we determined the most robust list of autophagosome proteins to date (Mancias et al., Nature 2014). Investigating pancreatic cancer specific autophagosomal candidates may reveal insights into how pancreatic cancer utilizes autophagy. Candidates will be characterized using an integrated proteomic, biochemical, and cell biologic approach. Furthermore, as we know little about the autophagosome machinery for selective autophagy (mitophagy, etc.), we will perform quantitative autophagosomal proteomics after various selective autophagy stimuli. These studies will yield further insights into autophagy biology (Mancias et al., JMB 2016). |
Identifying key KRas regulatory networks in Pancreatic Cancer
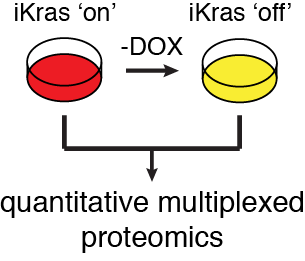
As KRas mutation is a hallmark of pancreatic cancer, understanding how oncogenic KRas signaling controls the proteome may lead to new therapeutic targets. Using tumors derived from a novel pancreatic cancer mouse model with an inducible oncogenic KRas allele in the pancreas (Ying et al., Cell 2012), quantitative whole cell and post-translational modification proteomics will be performed in order to characterize the global control of the proteome and signaling cascades by oncogenic KRas in vivo. Quantitative interaction proteomics -/+ oncogenic KRas will facilitate further functional dissection of pathways and interaction networks. Novel pathways will be explored using a combination of in vitro and in vivo pancreatic cancer tumor biology assays. Advanced quantitative proteomics promises to yield insights into pancreatic cancer biology and reveal new therapeutic targets. |
Identifying the pancreatic tumor MHC-I ligandome in response to ionizing radiation for combination radiation-immunotherapy approaches
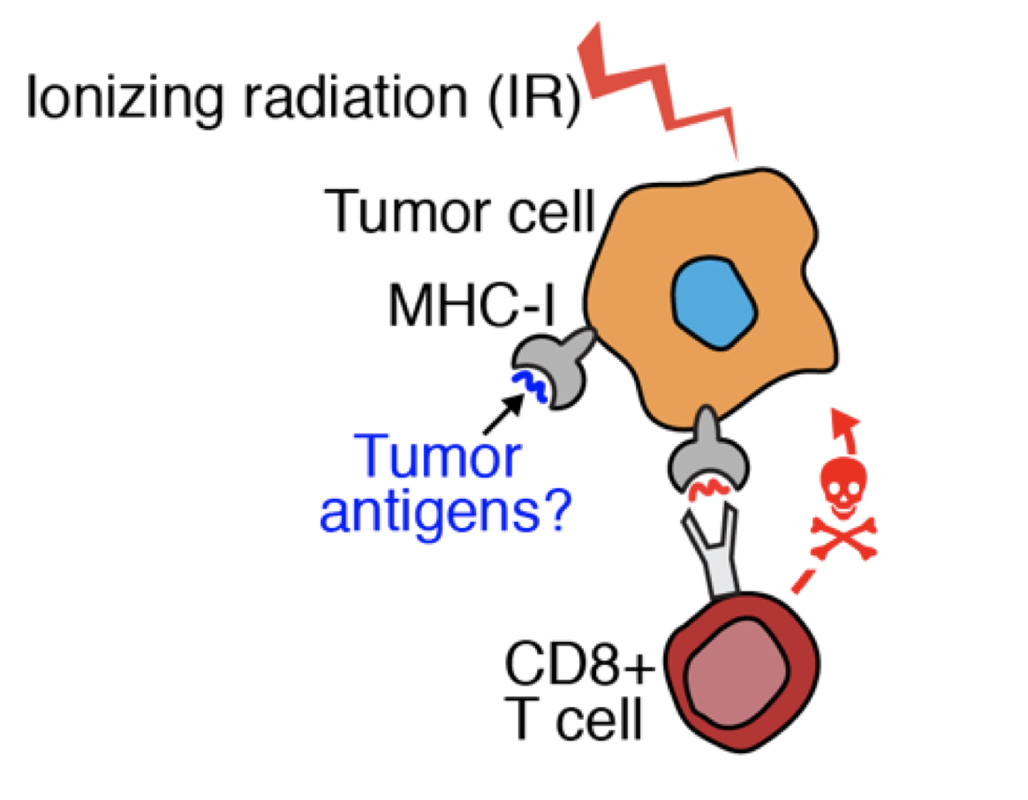
Radiation therapy has known stimulatory effects on anti-tumor CD8+ T cell immunity against both primary tumors and metastatic tumors distant from the primary irradiation site when combined with checkpoint blockade immunotherapy (abscopal effect). Here we aim to answer major questions regarding radiation-induced immune remodeling in pancreatic cancer: what are the MHC-I ligands pancreatic tumor cells present in response to ionizing radiation that stimulate CD8+ T cell mediated killing? To answer this question, we will use an innovative and cutting-edge combination of mass spectrometry-based quantitative peptidomics and proteomics, mouse models of PDAC, next-generation sequencing, bioinformatics, and targeted immune stimulation experiments. To that end, we have already developed an innovative multiplexed quantitative mass spectrometry-based approach towards identifying MHC-I ligands (Murphy, Yu et al, Analytical Chemistry 2019). |