
 Clara Poupault joins the lab today as a Research Technician! Clara just graduated from University of Massachusetts Boston with a BS in Biology (Honors). During her undergraduate studies, she worked for 3 years in the lab of Dr. Rister studying Vitamin A deprivation as a novel approach to identify neuroprotective molecules in a Drosophila model. Clara is interested in pursuing a MD-PhD program in the future. We're excited for Clara to begin working in the lab on projects related to radiation resistance and pancreatic cancer biology.
0 Comments
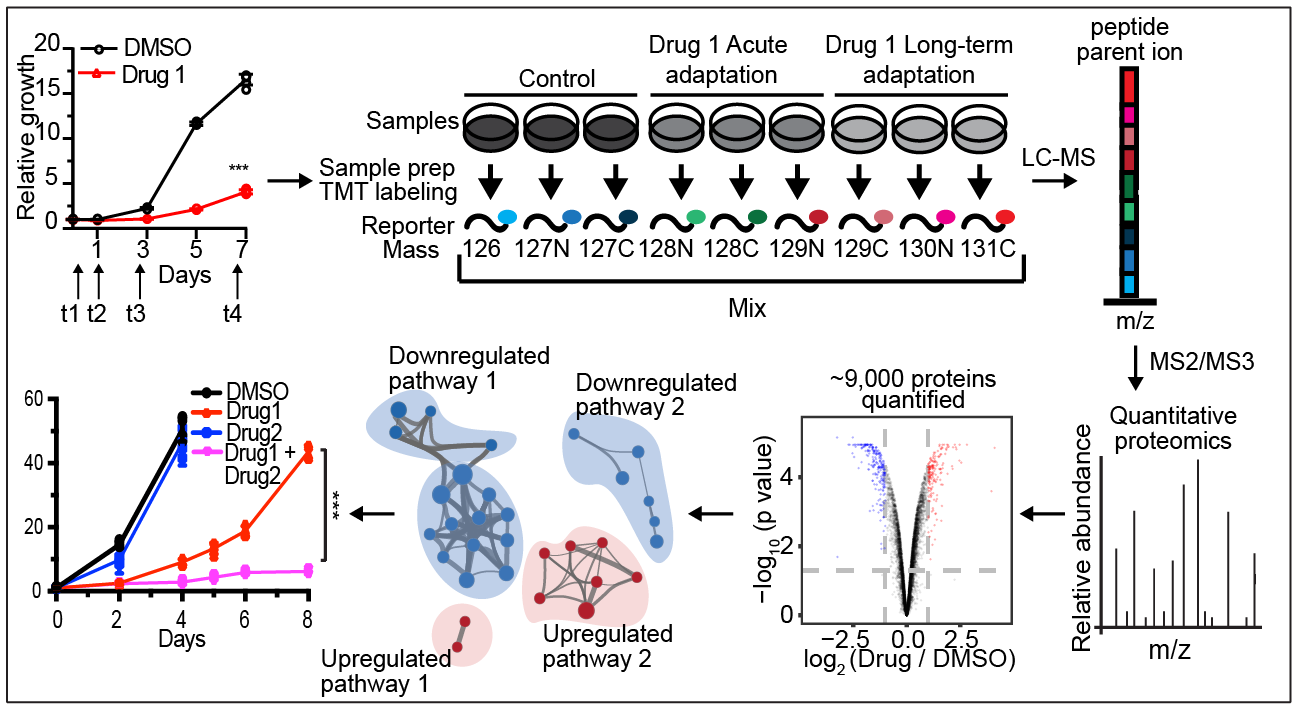
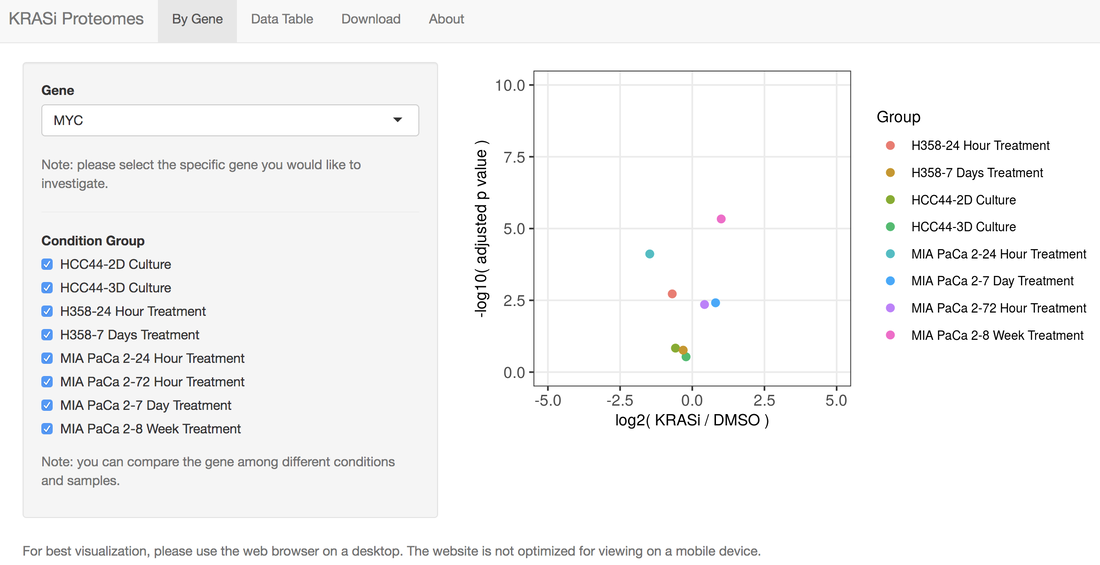
 Kristen John joins us as a research technician today! Welcome to the lab Kristen! Kristen just graduated from Hofstra University after majoring in Cell and Molecular Biology with a minor in Biochemistry. She previously worked in the lab of Jason Sheltzer, PhD at Cold Spring Harbor Laboratory using CRISPR/Cas9 to identify the correct molecular target of previously mischaracterized anti-cancer drugs and is a co-author on a paper published in Science Translational Medicine describing this work. We'er excited for Kristen to join us and help with projects in the lab on understanding resistance to radiation therapy using CRISPR/Cas9 screening. 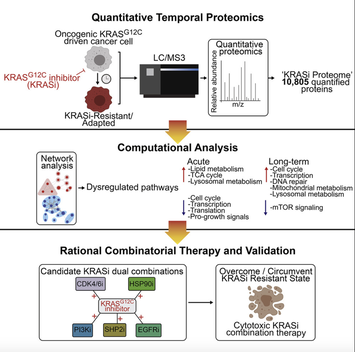 In this study published in Cell Reports and led by co-first authors Naiara Santana-Codina, PhD and Amrita Singh Chandhoke, PhD, the lab uses a quantitative temporal proteomics workflow to identify proteomic adaptations to short and long-term KRAS-G12C inhibition (KRASi) in lung and pancreatic adenocarcinoma. In this study, we quantified 10,805 proteins, representing the most comprehensive KRASi proteome (https://manciaslab.shinyapps.io/KRASi/). Our data reveal common mechanisms of acute and long-term response between KRASG12C-driven tumors. Based on these proteomic data, we identify potent combinations of KRASi with phosphatidylinositol 3-kinase (PI3K), HSP90, CDK4/6, and SHP2 inhibitors, in some instances converting a cytostatic response to KRASi monotherapy to a cytotoxic response to combination treatment. Overall, using quantitative temporal proteomics, we comprehensively characterize adaptations to KRASi and identify combinatorial regimens with potential therapeutic utility.
A funded postdoctoral fellow position is available in our lab. This is a great opportunity to study pancreatic cancer biology in our group. We have ongoing projects in: selective autophagy, therapeutic resistance, and radiation-induced immune responses. Please see the link for the full job description: https://careers-dfci.icims.com/jobs/18785/research-fellow/job?mode=view&mobile=false&width=783&height=500&bga=true&needsRedirect=false&jan1offset=-300&jun1offset=-240
|
Archives
September 2022
Categories |





 RSS Feed
RSS Feed